 
Hsp40 delivers substrates to the open, ATP-bound state of Hsp70 (Fig. The ATP-driven conformational cycle of Hsp70 is coordinated by Hsp40-class J-domain proteins (DnaJ in bacteria) and nucleotide exchange factors (NEFs GrpE in bacteria) ]. Reversible binding of hydrophobic peptides to the C-terminal substrate-binding domain (SBD) of Hsp70 is regulated by ATP binding and hydrolysis at the nucleotide-binding domain (NBD) (Fig. These diverse activities exploit the affinity of Hsp70 for short (5–7 amino acid) sequence elements enriched in hydrophobic residues (often flanked by positively charged amino acids) ] that are typically exposed by proteins in non-native conformations. Catalysis of folding by the Hsp70 chaperone systemĬhaperones of the Hsp70 class (DnaK in bacteria) are highly allosteric molecular machines that participate in a range of cellular processes, including protein folding and refolding, trafficking, translocation, disaggregation, and degradation ]. Here, we discuss examples of folding catalysis by the ATP-dependent Hsp70 and chaperonin (Hsp60) class of chaperones, and explore possible underlying mechanisms. Recent research has advanced the idea that molecular chaperones can modulate folding energy landscapes. This activity serves to inhibit aggregation, but can also influence the intramolecular interactions that define a protein's folding pathway. Chaperone classes use variations of a common mechanism of action based on transient binding of sequences enriched in hydrophobic residues. Molecular chaperones have evolved in response to these challenges and have in turn contributed to the diversity of proteomes in both prokaryotes and eukaryotes ]. Moreover, protein folding occurs in the context of translation ], which entails that nascent polypeptides are exposed to the cellular environment in an incomplete state lacking structural information needed for stable folding ]. The high concentration of macromolecules in the cell enhances the tendency of non-native proteins to aggregate ], while proteotoxic stress destabilizes the native state. The folding problem is exacerbated by conditions in vivo. The intrinsic challenges associated with folding of the larger, structurally more complex proteins that constitute the majority of proteomes ] may be more pronounced than current folding models suggest, with misfolding being the rule rather than the exception. Biophysical studies of protein folding typically focus on small model proteins (often < 100 amino acids) that are simple to express recombinantly and show robust reversible folding in vitro ]. Moreover, folding intermediates expose hydrophobic surfaces that can engage in nonfunctional intermolecular interactions enabling aggregation (Fig. Off-pathway intermediates and kinetic traps slow folding, and non-native intramolecular interactions can lead to stably misfolded states ]. 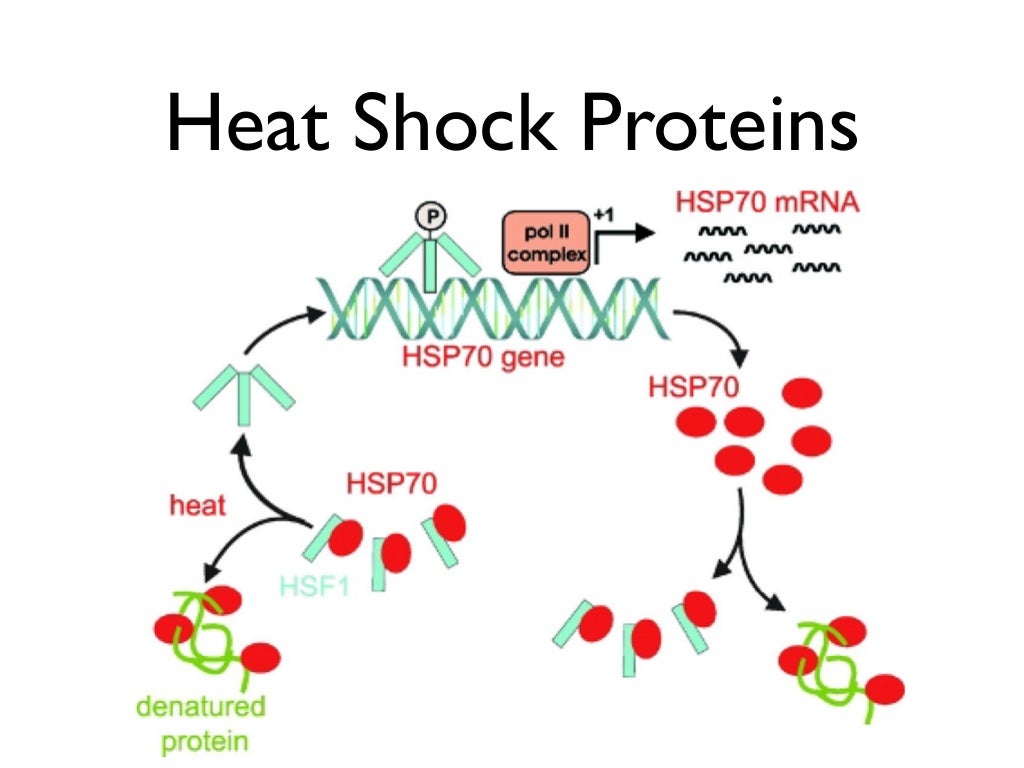
The folding free-energy landscape is rugged: Protein chains must traverse substantial energy barriers en route to the native state and consequently populate folding intermediates (Fig. Several factors complicate the folding process. Molecular chaperones inhibit aggregation, resolve kinetically trapped conformations, and provide kinetic assistance to folding by lowering free-energy barriers that separate folding intermediates from the native state. The accumulation of on- and off-pathway intermediates slows folding and entails the risk of misfolding into kinetically trapped states that are prone to form thermodynamically stable aggregates. 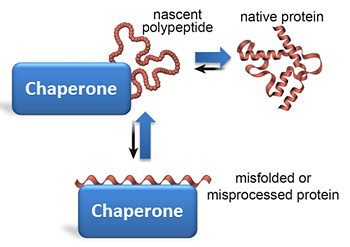
During folding, proteins navigate a rugged, funnel-shaped potential free-energy surface en route to the native state. Molecular chaperones shape the energy landscape of protein folding. Current models are not generally predictive for protein folding pathways, even if substantial progress has been made toward prediction of protein folds ]. However, a unifying mechanism for protein folding remains elusive ]. The conformational search for the native state is thought to follow a funnel-shaped energy landscape, driven by the burial of hydrophobic residues, and the relative stability of native-like interactions that nucleate the folding reaction ] (Fig. Subsequent work over the last 50 years has provided detailed insight into the general principles that govern protein folding. Pioneering experiments by Anfinsen in the 1950s ] demonstrated that a small protein can fold spontaneously in the absence of additional factors in vitro. H/DX-MS, hydrogen/deuterium exchange mass spectrometry 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
